Image enhancement
Partial volume effects correction
We investigated the factors that affect the accuracy of these corrections: segmentation of structures of interest from the anatomical MR data, spatial co-registration of the MR and PET volumes, characterization of the scanner’s point spread function (PSF) and the assumptions made during the correction (Bowen 2013).
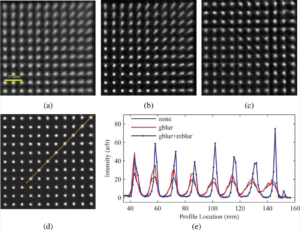
The influence of PSF based OSEM on 68Ge point source measurement image quality is shown in the figure below. Transaxial images showing a quadrant of the FOV reconstructed with (a) no PSF modelling, and modelling of the (b) spatially invariant Gaussian PSF (gblur) alone, (c) radial motion blurring alone (mblur), and (d) a combination of the Gaussian and radial motion blur kernels, and (e) line profile comparison along band in (d) (profile location of 0 is at the centre of the FOV).
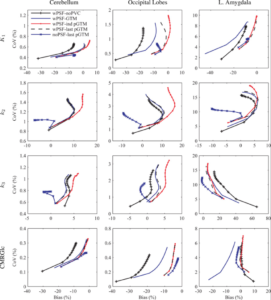
We explored the influence of the partial volume effects correction method and implementation on kinetic parameters estimated by fitting 18F-FDG dynamic data acquired on the BrainPET scanner and reconstructed with and without PSF modeling in the OSEM algorithm. Our results indicated that the partial volume effects correction implementation and choice of PSF modeling in the reconstruction can significantly impact model parameters (Bowen 2013).
Anatomy-aided PET image reconstruction
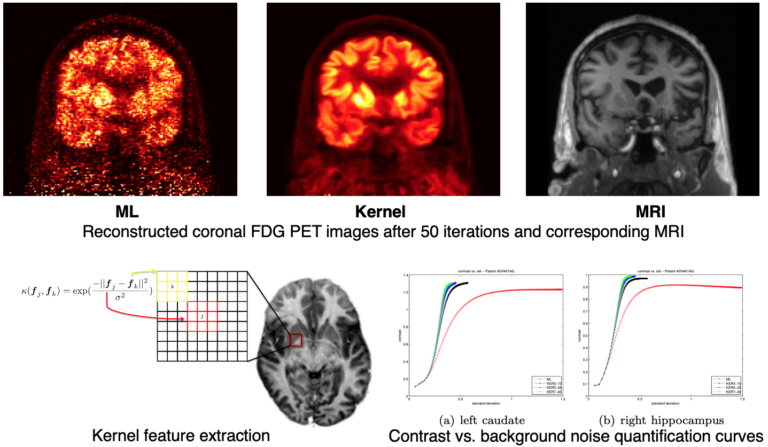
As a more elegant way to address this issue, in collaboration with Jinyi Qi’s group from the University of California at Davis, we have incorporated anatomical priors derived from high-resolution MRI into the PET image reconstruction model (Hutchcroft 2016).
MR-assisted PET data optimization
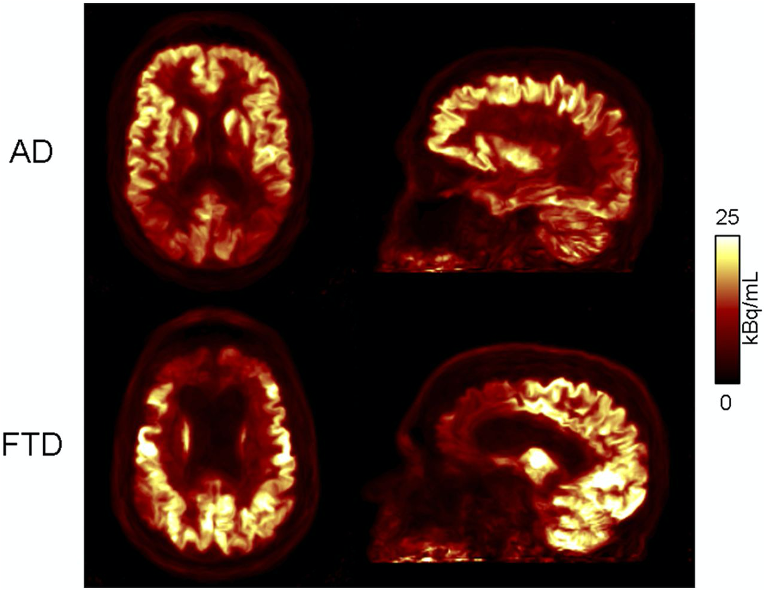
In addition to developing these individual methods, a major focus in our group has been on developing a unified data processing pipeline for integrating all these tools with the goal of improving the PET data quantification. We proposed an efficient algorithm to derive all the information required for performing: head attenuation and motion corrections, anatomy-aided reconstruction, and region-based analysis from the standard data acquired in ~6 minutes using a single morphological MR sequence with embedded motion navigators (Chen 2019).
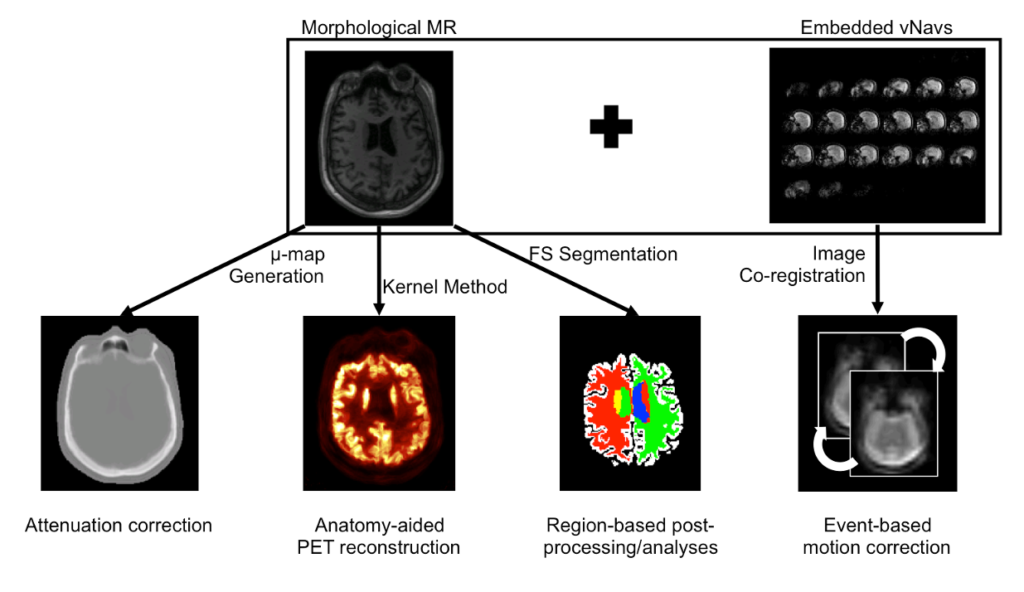
Using this approach, reduced variability and increased signal-to-noise ratio were seen after motion correction and anatomy-aided reconstruction. These results suggested PET data optimization may enable a more careful assessment of subtle changes in brain metabolism and allow for reduced sample sizes in future clinical trials.
Kinetic-guided image reconstruction
In the context of dynamic imaging, the conventional processing pipeline consists of independent image reconstruction of single time frames, followed by the application of a suitable kinetic model to each time series at the voxel or region-of-interest level. This approach ignores the existence of a temporal correlation between the signal value at each voxel in consecutive time frames, missing out on a powerful tool to solve the very under-determined inverse problem of image reconstruction.
Kinetic-guided PET reconstruction
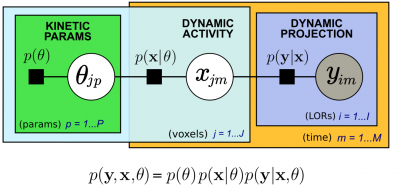
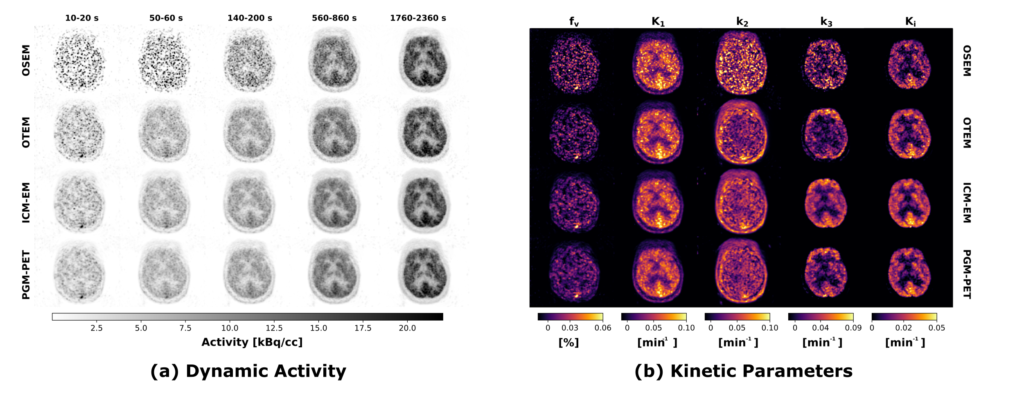
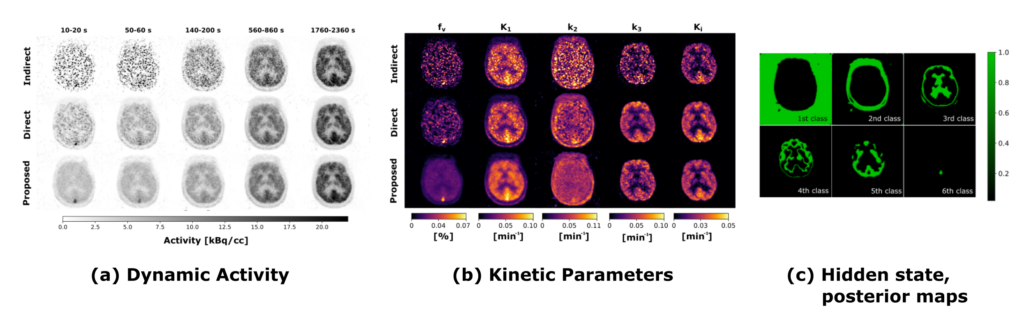
We introduced a new hierarchical model, which we formulate using the formalism of Probabilistic Graphical Modeling, in which kinetic modeling results are treated as prior expectation of activity time course, making it possible to control the trade-off between a data-driven and a model-driven reconstruction (Scipioni 2020), providing a bridge between them by properly tuning the impact of the kinetic modeling step on image reconstruction.
The proposed method is flexible to an arbitrary choice of (linear and nonlinear) kinetic models, it enables the inclusion of arbitrary (sub)differentiable priors for parametric maps, and it is simple to implement (Scipioni 2017 – IEEE NSS/MIC).
Kinetic-guided DCE-MRI reconstruction
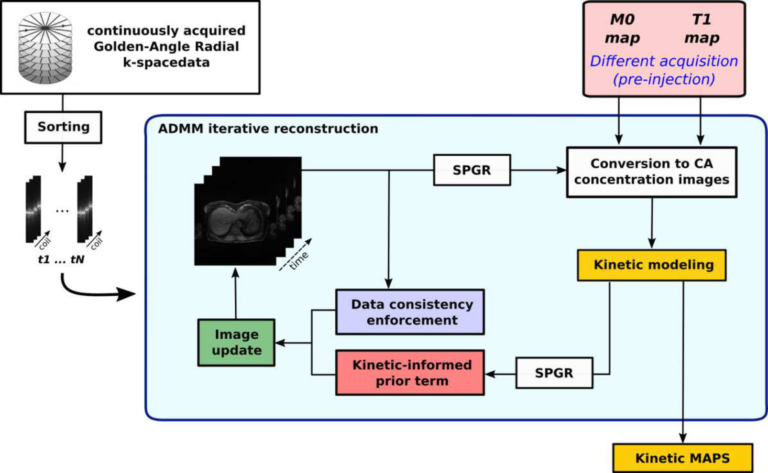
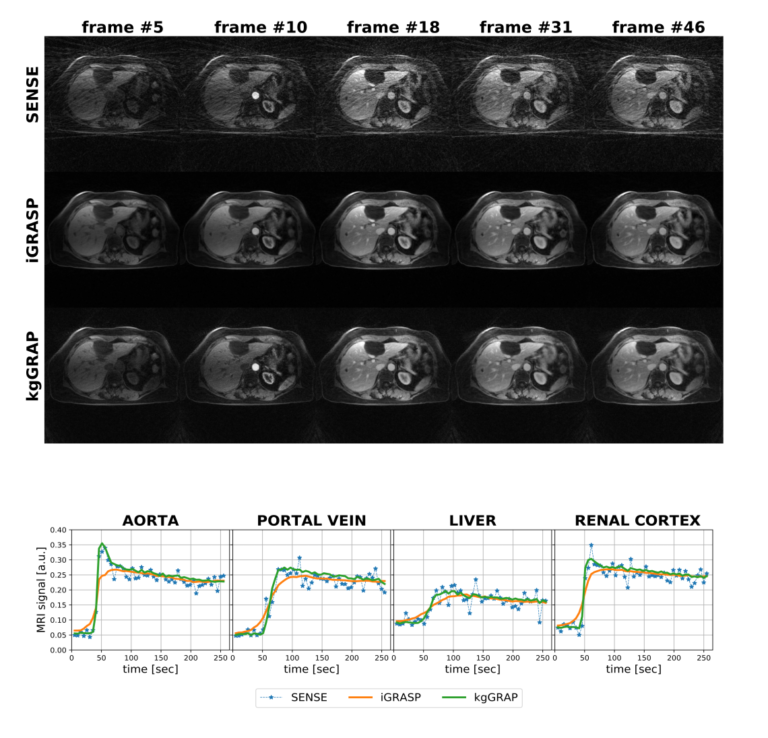
Building on the promising results obtained while working of 4D PET datasets, we decided to adapt the idea to obtain a reconstruction method for free-breathing DCE-MRI, that combines golden-angle radial sampling, parallel imaging and tracer kinetic (TK) modeling in an iterative reconstruction approach. Once again, information coming from TK modeling are treated as a priori knowledge, assisting image reconstruction. When compared with Total Variation Compressed Sensing reconstruction, this approach achieves comparable de-noising effect in the spatial domain, but improved temporal fidelity and TK modeling. (Scipioni 2019 – ISMRM19)
Direct 4D PET reconstruction with discrete tissue types
Trying to push the boundaries even further, we also evaluated a novel direct reconstruction framework, which include concurrent clustering as a potential aid in addressing high levels of noise typical of voxel-wise kinetic modeling. Core assumption is that the imaged volume is formed by a finite number of different functional regions, and that voxel-wise time courses are determined by the functional cluster they belong to. Probabilistic Graphical Modeling theory is used to describe the problem, and to derive the inference strategy. The proposed iterative estimation scheme provides concurrent estimate of kinetic parameter maps, activity images, and segmented clusters (Scipioni 2019 – EMBC).