Attenuation correction
Head
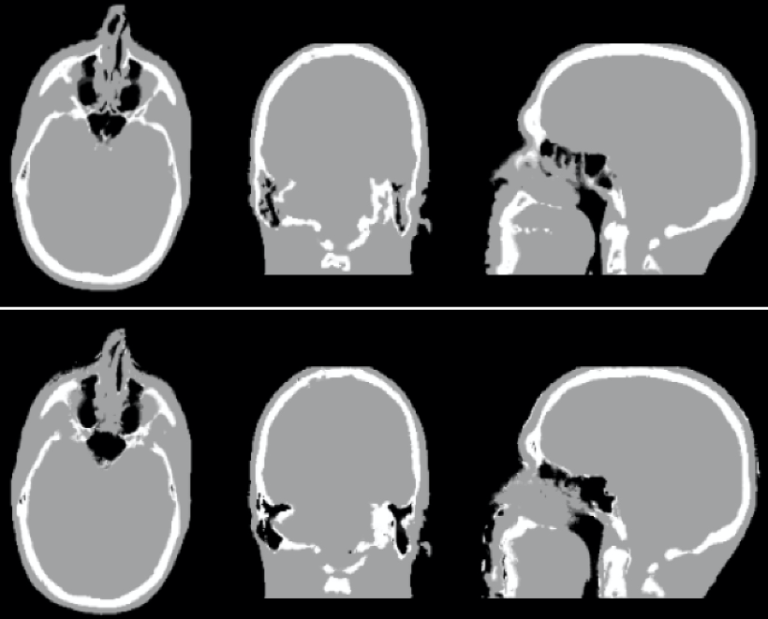
Identifying bone tissue is particularly relevant for accurate attenuation correction in neurological PET studies as this tissue class has the highest linear attenuation coefficient and inaccuracies in its estimation can introduce large biases in the adjacently located cortical structures. Since using conventional MRI pulse sequences bone tissue and air-filled cavities are difficult to distinguish, novel sequences have been developed to address this challenge. We were among the first to propose using ultra-short echo time MR sequences (optimized for imaging tissues with very short T2 relaxation times) for generating segmented head attenuation maps. In a first approach, the head was segmented into three compartments (i.e. bone and soft tissue and air cavities) based on the relationship between the two echoes on a voxel-by-voxel basis (Catana 2010).
Subsequently, combining the dual-echo UTE and T1-weighted MR data using probabilistic atlases allowed the generation of substantially improved segmented attenuation maps (Poynton, Chen et al 2014). However, one of the main limitations of early generation UTE-based methods was that only three compartments (i.e. soft tissue, bone, and air cavities) could be identified.
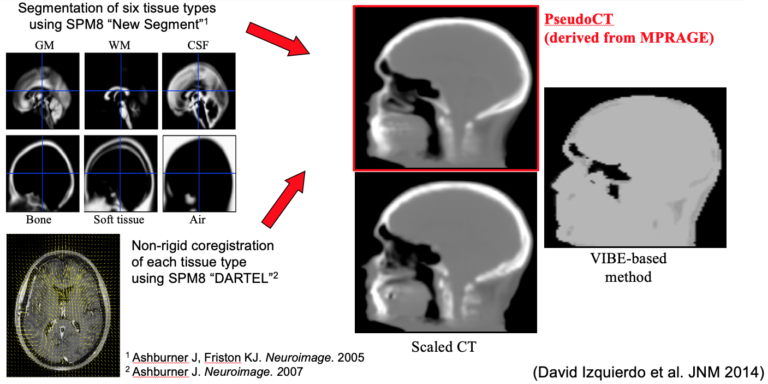
We next implemented an SPM-based method for generating head attenuation maps from a single morphological MRI dataset obtained with the MPRAGE sequence routinely collected in research neurological studies (Izquierdo-Garcia et al 2014). After intensity normalization, the MR images are segmented into six tissue classes using the “New Segment” SPM tool and then registered to a previously created template using a diffeomorphic non-rigid image registration algorithm (SPM DARTEL). The inverse transformation is applied to obtain the pseudo-CT images in the subject space. Using this approach, the voxel- and regional-based quantification errors compared to the scaled CT method were 3.87±5.0% and 2.74±2.28%, respectively. This method was also demonstrated to work in patients with modified anatomy (e.g. glioblastoma patients post-surgery). This method and several of the others developed by other groups around the world have been shown in a multi-center evaluation to be quantitatively accurate to a degree “smaller than the quantification reproducibility in PET imaging” (Ladefoged 2017). Continuous-valued attenuation maps were also obtained from dual-echo UTE and T1-weighted MR data using probabilistic atlases (Chen et al 2017). We also assessed the repeatability of the atlas- or UTE-based methods. For example, comparable attenuation maps and PET volumes were obtained at three visits using the probabilistic atlas method (Chen et al 2017). Similar results were reported for the SPM-based method (Izquierdo-Garcia et al 2018).
Pelvis
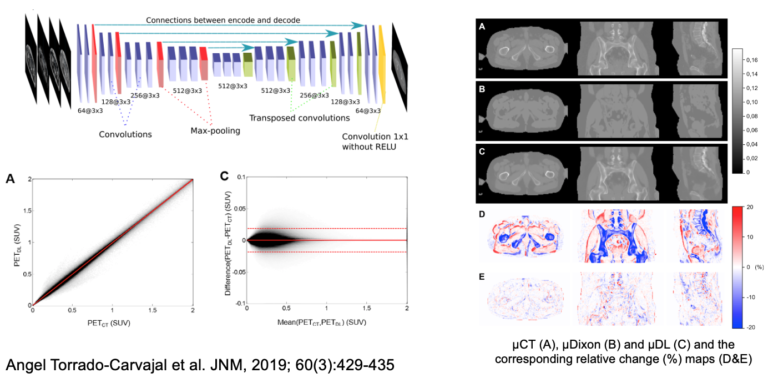
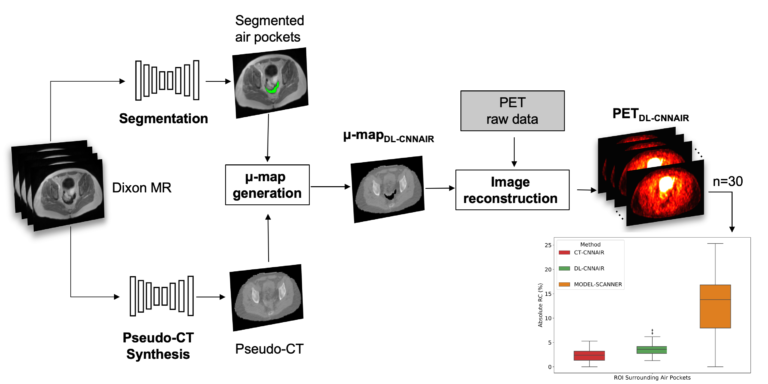
The pelvis is another area where MR-based approaches that do not properly account for bone attenuation could lead to substantial bias in PET data quantification and deep learning approaches have been proposed to minimize this bias starting from the MR data acquired using specialized MR sequences. Instead, we implemented a deep learning-based approach to generate pseudo-CT maps exclusively from the Dixon-VIBE MR images routinely acquired for attenuation correction on the Biograph mMR scanner. Using our 2D network that takes four contrasts as inputs (water, fat, in-phase, and out-of-phase Dixon-VIBE images), the mean absolute relative change in PET values in the pelvis area was 2.36% ± 3.15% (Torrado-Carvajal et al 2019).
One remaining challenge in using convolutional neuronal networks to synthesize CT from MR images of the pelvis is the presence of air pockets (i.e., digestive tract gas) in this area. As CT and MR images are acquired on separate scanners at different times, the locations and sizes of these air pockets can change between the two scans, which can lead to errors in both the MR-CT co-registration and image synthesis tasks. We trained and evaluated CNNs to automatically segment air pockets from MR CAIPIRINHA-accelerated Dixon images and assessed the quantitative impact on the reconstructed PET images (Sari et al, submitted to JNM).
Metallic implants
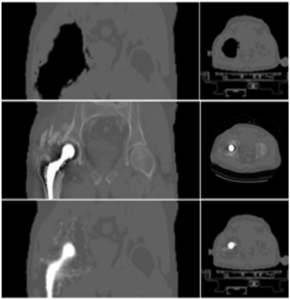
The numerous foreign objects (e.g. dental implants, surgical clips and wires, orthopedic screws and plates, prosthetic devices, etc.) that can be present in the subject lead to susceptibility artifacts in the MR images that propagate as signal voids in the corresponding attenuation maps.
We developed a method to estimate the location, shape, and linear attenuation coefficient of the implant using a joint reconstruction of the activity and attenuation algorithm. The implant PET-based attenuation map completion (IPAC) method performs a join reconstruction of radioactivity and attenuation from the emission data (Fuin et al 2017).
Transmission imaging on the Biograph mMR
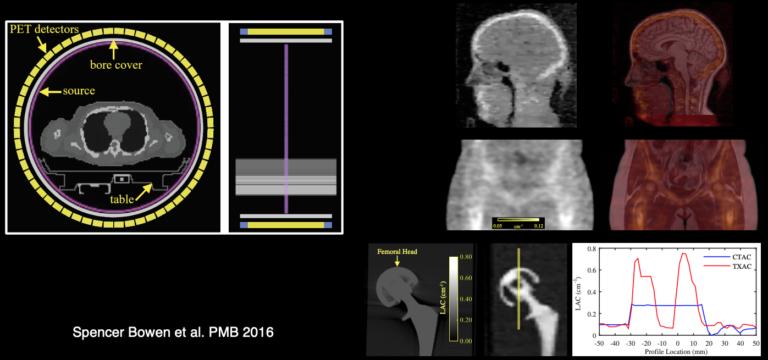
While transmission-based techniques are still considered the true gold standard for PET attenuation correction, traditional rotating transmission sources have not been integrated into the PET/MRI scanners due to obvious engineering challenges and the desire to reduce the radiation exposure. As such techniques would be valuable for improving and validating MR-based approaches, several stationary transmission sources have been suggested as alternatives.
In the case of scanners without time-of-flight capabilities (e.g. Biograph mMR), we showed that a single torus source filled with [18F]FDG allows the acquisition of highly accurate transmission images before radiotracer administration (Bowen et al 2016).