our research
SOFTWARE
Develop methods to synergistically use the data.
Attenuation
Correction
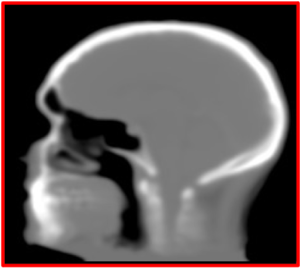
The methods traditionally used for accounting for the absorption of the 511 keV annihilation photons before they reach the PET detectors (by interactions with the subject and other materials placed in the PET field of view) are not available in integrated PET/MRI scanners and approaches for deriving similar information from the MR images had to be developed.
Several aspects must be considered when implementing an MR-based attenuation correction method. The MR signal intensity, which depends on proton density and tissue relaxation times, does not directly reflect electron density, which is relevant for attenuation correction. This makes bone tissue difficult to visualize using the data acquired using conventional MR sequences. Similarly, MR imaging of lung tissue is challenging given its low proton density, susceptibility artifacts at the many air-tissue interfaces, tissue fraction effects, respiratory motion, etc. Subject motion leads to mismatches between the MR and the emission data. The patient’s arms are positioned along the body for PET/MRI studies and thus extend outside the uniform region of the MR field-of-view, leading to truncation artifacts. Foreign objects (e.g. dental implants and prostheses) can cause susceptibility artifacts in the MR images. Intravenous contrast agents often used for clinical purposes can change the contrast between different structures (Catana 2020).
People
Research areas:
Motion
Correction
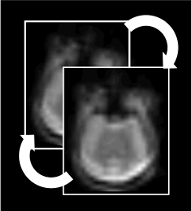
Simultaneous PET and MRI data acquisition immediately brings to mind the possibility of improving the performance and information content of one instrument using the information obtained from the other modality. One such example, explored in our laboratory, is MR-assisted motion correction. PET studies are usually long and subject motion (voluntary and involuntary) is difficult to avoid, leading to degradation (blurring) of the images and to severe artifacts when motion amplitude is large, often offsetting the benefit of using a high-resolution scanner (Catana 2015).
People
Research areas:
Image
Enhancement
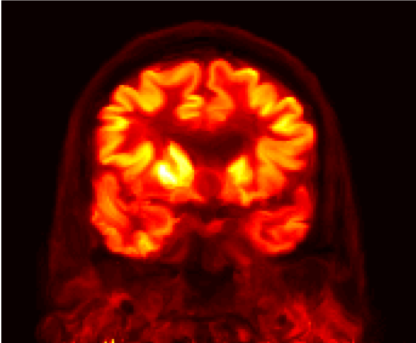
Partial volume effects lead in PET to an under-or overestimation of tissue activity concentrations that depends on the activity distribution and the size and shape of the structures of interest, virtually affecting all neurological studies performed today. Although a myriad of partial volume effect correction methods has previously been proposed, no one method has been accepted or is routinely used for research applications and even less so in the clinical practice. We investigated the factors that affect the accuracy of these corrections: segmentation of structures of interest from the anatomical MR data, spatial co-registration of the MR and PET volumes, characterization of the scanner’s point spread function and the assumptions made during the correction (Bowen 2013).
As a more elegant way to address this issue, in collaboration with Jinyi Qi’s group from the University of California at Davis, we have incorporated anatomical priors derived from high-resolution MRI into the PET image reconstruction model (Hutchcroft 2016).
In addition to developing these individual methods, a major focus in our group has been on developing a unified data processing pipeline for integrating all these tools with the goal of improving the PET data quantification. We proposed an efficient algorithm to derive all the information required for performing: head attenuation and motion corrections, anatomy-aided reconstruction, and region-based analysis from the standard data acquired in ~6 minutes using a single morphological MR sequence with embedded motion navigators (Chen 2019).
Using this approach, reduced variability and increased signal-to-noise ratio were seen after motion correction and anatomy-aided reconstruction. These results suggested PET data optimization may enable a more careful assessment of subtle changes in brain metabolism and allow for reduced sample sizes in future clinical trials.
People
Research areas:
Kinetic
Modeling
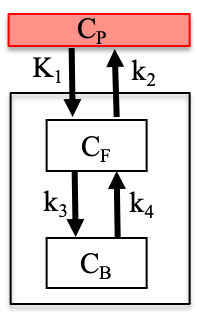
Kinetic modeling of dynamically acquired time series is necessary to achieve full quantitation of PET or DCE-MRI imaging data. Sets of differential equations can be used to describe the changing tracer/contrast concentrations in biologically distinct compartments (i.e. blood plasma, free in tissue, bound to receptors). Which specific kinetic model is used depends on the radiotracer/contrast agent used, the study design, and the tissue of interest.
People
Research areas:
PET kinetic modeling
Kinetic-guided image reconstruction
Machine
Learning
Artificial intelligence is increasingly used for a range of applications in healthcare, including medical imaging. We have also embraced these promising techniques and developed machine learning techniques for several applications.
People
Research areas:
Attenuation correction for pelvis PET-MR imaging
MR-based motion estimation for cardiac PET-MRI studies
Myocardial strain estimation using DeepStrain
Multimodal primary prostate cancer characterization